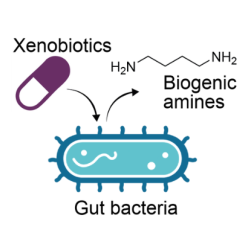
Microbiome mediated toxicity of drugs and other xenobiotics
Prof. Kiran Patil
We are investigating microbiome-drug and microbiome-pollutant interactions to uncover their contribution to side effects and toxicity.
An adult individual typically harbours a few hundred grams of bacteria in their digestive tract. Recent findings have brought forward the fundamental role of this microbiota in determining drug efficacy, mode of action and side effects. Yet, such interactions at the microbiome level remain unknown for the vast majority of the drugs and for other xenobiotics that we are exposed to (e.g. food contaminants like pesticides). The overarching goal of Kiran’s research programme is to discover and model complex (xeno-) metabolic networks emerging in the gut microbiota and thereby gain mechanistic insights into microbiome-mediated toxicity. To this end, this interdisciplinary programme brings together computational (metabolic modelling, bioinformatics, chemo-informatics and machine learning) and experimental (in vitro microbiomics, metabolomics and chemical genomics) approaches.
Key publications
Lindell, A.E., Grießhammer, A., Michaelis, Papagiannidis, D., Ochner, H., Kamrad, S., Guan, R., Blasche, S., Ventimiglia, L.N., Ramachandran, B., Ozgur, H., Zelezniak, A., Beristain-Covarrubias, N., Yam-Puc, J. C., Roux, I., Barron, L. P., Richardson, A.K., Guerra Martin, M., Benes, V., Morone, N., J.E.D, Bharat,T.A.M., Savitski, M.M., Maier, L., & Patil, K.R., Human gut bacteria bioaccumulate per- and polyfluoroalkyl substances. Nat Microbiol 10, 1630–1647 (2025). https://doi.org/10.1038/s41564-025-02032-5
Lindell AE, Zimmermann-Kogadeeva M and Patil KR. Multimodal interactions of drugs, natural compounds and pollutants with the gut microbiota. Nat Rev Microbiol, doi:10.1038/s41579-022-00681-5 (2022)
Klunemann M, Andrejev S, Blasche S, Mateus A, Phapale P, Devendran S, Vappiani J, Simon B, Scott TA, Kafkia E, Konstantinidis D, Zirngibl K, Mastrorilli E, Banzhaf M, Mackmull MT, Hovelmann F, Nesme L, Brochado AR, Maier L, Bock T, Periwal V, Kumar M, Kim Y, Tramontano M, Schultz C, Beck M, Hennig J, Zimmermann M, Sevin DC, Cabreiro F, Savitski MM, Bork P, Typas A and Patil KR. Bioaccumulation of therapeutic drugs by human gut bacteria. Nature 597(7877): 533-538 (2021)
Machado D, Maistrenko OM, Andrejev S, Kim Y, Bork P, Patil KR and Patil KR. Polarization of microbial communities between competitive and cooperative metabolism. Nat Ecol Evol 5, 195–203 (2021)
Maier L, Pruteanu M, Kuhn M, Zeller G, Telzerow A, Anderson EE, Brochado AR, Fernandez KC, Dose H, Mori H, Patil KR, Bork P and Typas A. Extensive impact of non-antibiotic drugs on human gut bacteria. Nature 555, 623–628 (2018)
Tramontano M, Andrejev S, Pruteanu M, Klünemann M, Kuhn M, Galardini M, Jouhten P, Zelezniak A, Zeller G, Bork P, Typas A and Patil KR. Nutritional preferences of human gut bacteria reveal their metabolic idiosyncrasies. Nature Microbiol. 3, 514–522 (2018)